 » Barrier-free View
» Barrier-free View
| ||||||||
 | ||||||||
 |
 |
 |
| Picture of the 1st Network Meeting of the Herz-Kreislauf-NetzDownloadsArchivExternal job opportunities |
 |
Identification of genetic modifiers of cardiomyopathies in mice
 |
Subproject leader
Dr. Boris Ivandic
University of Heidelberg
Internal Medicine III / Cardiology
Otto-Meyerhof-Center
Im Neuenheimer Feld 350
69120 Heidelberg
email: boris.ivandic@med.uni-heidelberg.de
Phone: 06221-56 8853
Systematic investigation of the influence of modifier loci on cardiomyopathy phenotypes is difficult in humans, because environmental factors cannot be controlled and genetic variation is large. Association studies manage this problem either by substantially increasing the numbers of cases and controls or by studying exclusively familial cases of cardiomyopathy. In rodent models, however, environmental and genetic complexity can be reduced much further: mouse inbred strains allow to examine multiple animals with a defined genetic background in a controlled environment. Employing quantitative trait locus analysis (QTL) we identified previously genetic modifiers conferring susceptibility to calcification of dystrophic or necrotic myocardium. In mice, this calcifying phenotype of cardiomyopathy is also referred to as dystrophic cardiac calcification. To simplify the identification of loci modifying a disease phenotype one can use a so-called disease-sensitized mouse model carrying a mutant disease-causing allele on a susceptible, inbred background. Recently, such congenic mutant strains were used to map modifier loci of cardiomyopathy by QTL analysis. The underlying genes, however, remain to be determined by positional cloning, which may take years of work.
As a faster alternative, we apply now micro array technology to identify disease-specific and genetic modifier pathways. For this purpose, we will generate congenic strains transferring the genetic defects from several established cardiomyopathy models (on a mixed C57BL/6 x 129 background) onto 4 inbred strain backgrounds (C57BL/6, C3H/He, FVB/N, 129S1/Sv).
The first specific aim is the identification of inbred strain backgrounds determining high and low disease penetrance by comprehensive characterisation of cardiac pathology and function. The second specific aim is the identification of pathways associated with either the disease-causing allele or the influence of modifier genes. This will be achieved by comparison of myocardial gene expression in strains sharing a mutant allele with highest and lowest disease penetrance.
Generation of mutant congenic strains. The novel mutant congenic strains are generated, phenotyped and analysed in close collaboration with the group of Prof. M. Hrabé de Angelis (SMP-Models, GSF Research Institute). Strains will be cryopreserved at the European Mouse Mutant Archive (EMMA). Mutant congenic strains are produced employing accelerated breeding protocols for so-called speed congenics: male pups are genotyped using SNP assays to apply negative and positive selection criteria (ie. small amounts of donor genome and heterozygosity of the mutant allele, respectively) in order to select suitable animals for further backcrossing to obtain the next generation (Fig. 1).
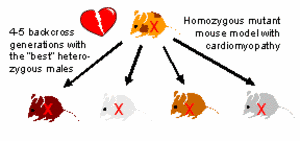 |
Fig. 1:. Generation of congenic mutant strains by breeding. |
Phenotyping. Phenotyping will be performed in the cardiovascular screening unit at the German Mouse Clinic (GMC) to determine the penetrance of the mutant allele (Fig. 2). Cardiomyopathy is characterised by echocardiography to assess systolic and diastolic ventricular dysfunction measured exactly by fractional shortening coefficients, E/A ratio and other parameters. Digital electrocardiography is performed in a highly standardized fashion to detect ECG abnormalities and arrhythmias. Treadmill stress tests are used to assess functional cardiovascular capacity. Morphological changes of the myocardium are examined by standard histology, immunohistochemistry and electron microscopy.
 |
Fig. 2: Extensive cardiac phenotyping to identify strains with high and low penetrance of the mutant allele. |
Microarray analysis. In cooperation with Dr. J. Beckers (MXpress facility, GSF Research Institute), genome-wide gene expression profiling will be assessed in congenic mutant strains, which share a mutant allele, but represent opposite extremes of the phenotype, i.e. exhibit the highest and the lowest penetrance of cardiomyopathy (Fig. 3). Bioinformatics software from Genomatix Inc. (Muenchen, Germany) will be used to assemble pathways of co-regulated genes: with respect to the selected strains, coordinate gene regulation is related to the mutant allele, while differential gene expression is interpreted as the signature of genetic modifiers in the respective strain backgrounds.
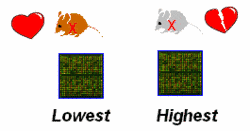 |
Fig. 3: Comparison of genome-wide gene expression in the strains, which exhibited the highest and the lowest penetrance of cardiomyopathy due to a shared mutant allele. |
Initially, the required numbers of animals of each cardiomyopathy mouse model (homozygous animals) were difficult to obtain, because the fertility of some mutant lines was significantly reduced owing to the severity of the cardiomyopathy phenotype (heart failure). At present, the mutant alleles of all cardiomyopathy models are transferred onto the inbred C57BL/6 background by repeated backcrossing for at least 4 generations. At the same time, transferral the mutant alleles onto the remaining strain backgrounds has begun. Phenotyping and micro array analyses will be started in the final year of NGFN.
This project is part of a scientific collaboration of the Cardiovascular Disease network with the SMP-Models. The mutant congenic strains are a valuable ressource for the identification of genetic modifiers of cardiomyopathy complementing association studies in DCM patients (see Project Contribution of Genetic Variation in Modifier Genes to Cardiomyopathy Phenotypes). Differentially regulated genes may point to novel candidate genes, which will be examined in other NGFN projects as well. The mutant congenic strains will be offered to collaborating scientists for detailed studies of cardiac physiology, myocardial cell biology and pharmacological intervention.
 |
Mutant mice |
 |
Working group Dr. Ivandic and PD Dr. Weichenhan |
Publications:
1. Ivandic BT, Utz HF, Kaczmarek PM, Aherrahrou Z, Axtner SB, Klepsch C, Lusis AJ, Katus HA. New Dyscalc loci for myocardial cell necrosis and calcification (dystrophic cardiac calcinosis) in mice. Physiol Genomics. 2001;6:137-44.
2. Ivandic BT, Qiao JH, Machleder D, Liao F, Drake TA, Lusis AJ. A locus on chromosome 7 determines myocardial cell necrosis and calcification (dystrophic cardiac calcinosis) in mice. Proc Natl Acad Sci U S A. 1996;93:5483-8.
3. Korff S, Riechert N, Schoensiegel F, Weichenhan D, Autschbach F, Katus HA, Ivandic BT. Calcification of myocardial necrosis is common in mice. Virchows Arch. 2005 Oct 7;:1-9 [Epub ahead of print] PMID: 16211391 [PubMed - as supplied by publisher]
4. Churchill GA, Airey DC, Allayee H, Angel JM, Attie AD, Beatty J, Beavis WD, Belknap JK, Bennett B, Berrettini W, Bleich A, Bogue M, Broman KW, Buck KJ, Buckler E, Burmeister M, Chesler EJ, Cheverud JM, Clapcote S, Cook MN, Cox RD, Crabbe JC, Crusio WE, Darvasi A, Deschepper CF, Doerge RW, Farber CR, Forejt J, Gaile D, Garlow SJ, Geiger H, Gershenfeld H, Gordon T, Gu J, Gu W, de Haan G, Hayes NL, Heller C, Himmelbauer H, Hitzemann R, Hunter K, Hsu HC, Iraqi FA, Ivandic B, Jacob HJ, Jansen RC, Jepsen KJ, Johnson DK, Johnson TE, Kempermann G, Kendziorski C, Kotb M, Kooy RF, Llamas B, Lammert F, Lassalle JM, Lowenstein PR, Lu L, Lusis A, Manly KF, Marcucio R, Matthews D, Medrano JF, Miller DR, Mittleman G, Mock BA, Mogil JS, Montagutelli X, Morahan G, Morris DG, Mott R, Nadeau JH, Nagase H, Nowakowski RS, O'Hara BF, Osadchuk AV, Page GP, Paigen B, Paigen K, Palmer AA, Pan HJ, Peltonen-Palotie L, Peirce J, Pomp D, Pravenec M, Prows DR, Qi Z, Reeves RH, Roder J, Rosen GD, Schadt EE, Schalkwyk LC, Seltzer Z, Shimomura K, Shou S, Sillanpaa MJ, Siracusa LD, Snoeck HW, Spearow JL, Svenson K, Tarantino LM, Threadgill D, Toth LA, Valdar W, de Villena FP, Warden C, Whatley S, Williams RW, Wiltshire T, Yi N, Zhang D, Zhang M, Zou F; Complex Trait Consortium. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat Genet. 2004 Nov;36(11):1133-7. PMID: 15514660 [PubMed - indexed for MEDLINE]
5. Ivandic BT, Qiao JH, Machleder D, Liao F, Drake TA, Lusis AJ. A locus on chromosome 7 determines myocardial cell necrosis and calcification (dystrophic cardiac calcinosis) in mice. Proc Natl Acad Sci U S A. 1996;93:5483-8.
6 .Aherrahrou Z, Axtner SB, Kaczmarek PM, Jurat A, Korff S, Doehring LC, Weichenhan D, Katus HA, Ivandic BT. A locus on chromosome 7 determines dramatic upregulation of osteopontin in dystrophic cardiac calcification in mice. Am J Pathol. 2004, 164:1379-87.
7. Matin A, Nadeau JH. Sensitized polygenic trait analysis. Trends Genet. 2001;17:727-31.
8. Montagutelli X. Effect of the genetic background on the phenotype of mouse mutations. J Am Soc Nephrol. 2000;11:S101-5.
9. Le Corvoisier P, Park HY, Carlson KM, Marchuk DA, Rockman HA. Multiple Quantitative Trait Loci Modify the Heart Failure Phenotype in Murine Cardiomyopathy. Hum Mol Genet. 2003;30:30.
10. Suzuki M, Carlson KM, Marchuk DA, Rockman HA. Genetic modifier loci affecting survival and cardiac function in murine dilated cardiomyopathy. Circulation. 2002;105:1824-9.
11. Wong GT. Speed congenics: applications for transgenic and knock-out mouse strains. Neuropeptides. 2002;36:230-
more Information about the working group:
http://www.klinikum.uni-heidelberg.de/index.php?id=4573