 » Barrier-free View
» Barrier-free View
| ||||||||
 | ||||||||
 |
 |
 |
| Picture of the 1st Network Meeting of the Herz-Kreislauf-NetzDownloadsArchivExternal job opportunities |
 |
Comparative genomic of left ventricular hypertrophy and dysfunction in hypertension
 |
Subproject leader
Prof. Dr. Reinhold Kreutz
Charite - Universitätsmedizin Berlin / Campus Benjamin Franklin
Institute of Clinical Pharmacology and Toxicology
Department Clinical Pharmacology
Hindenburgdamm 30
12200 Berlin
email: reinhold.kreutz@charite.de
Phone: 030-8445 2279
The aim of this project is the identification and characterization of new candidate genes and molecular pathways that are involved in the development of left ventricular (LV) hypertrophy (LVH) and LV dysfunction. Our research program in this project is carried out in two complementary topics.
Topic 1:
Comparative genomics to elucidate positional candidate regions for LVH.
A major goal of this subproject (topic 1) is to characterize genetic susceptibility loci that determine LV mass and thus LVH in hypertension. To this end we will follow a comparative genome approach to elucidate positional candidate regions for LVH by investigating candidate loci that have been identified as quantitative trait loci (QTL) for LV mass in inbred polygenetic rat models in human populations. These populations are available in KGCV via the network initiative on population genetics of cardiovascular disease (Schunkert, Kääb, Wichmann) and at the University of Nürnberg-Erlangen.
Two major QTL with a striking effect on LV mass were identified on rat chromosome 1 (RNO1, Figure 1) and on RNO19 (Siegel et al. 2003). At the QTL on RNO1 one allele of the stroke-prone spontaneously hypertensive (SHRSP) rat accounted for a 10% increase in LV mass (i.e. 20% increase in LV mass in homozygous animals, Figure 2), while this QTL was not linked to blood pressure (Figure 1).
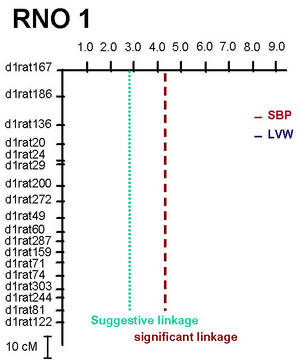 |
Fig 1: Lod-plot for systolic blood pressure (SBP) and left ventricular weight (LVW) on rat chromosome 1 (RNO1) in an F2-cross (n=232 male animals) between SHRSP and F344 rat strains. |
This mapping result was obtained by contrasting SHRSP with an inbred strain for low heart weight, i.e. the F344 rat. Our experimental strategy in the rat will focus on the characterization of two speed congenic strains for the QTLs on RNO1 and RNO19. Our comparative mapping strategy will include both candidate regions, while association studies in humans will at first focus on the candidate region on RNO1.
The comparative genome analysis will involve prediction of homologous genomic regions in the human genome. To this end we will follow a two-step genotyping approach in the MONICA population from Augsburg (Schunkert et al. 1994) involving currently 1600 individuals with LV mass phenotypes and functional data obtained by echocardiography. We will first perform genotyping in a subset of informative individuals (~400 individuals with high and ~400 individuals with low LV mass) with the entire SNP set in the candidate region (350 SNPs) followed by verification of positively associated SNPs in the whole MONICA study population. i.e. in 1600 individuals. Subsequently, we will perform detailed functional clinical testing of identified candidate genes in animal models and in clinical studies (biochemical analysis, physiologic and pharmacologic testing) in Erlangen in a very homogenous population of subjects.
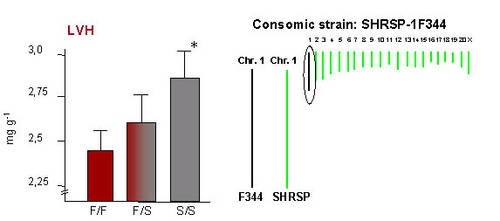 |
Fig 2: Left panel: Genotype-phenotype relation at the QTL for LVH (given as relative LV weight in mg/g) on RNO1. S, SHRSP allele; F, F344 allele. * p=1,7 * 10-9 vs. F/F. Right panle:The generation of the consomic strain SHSRP-1F344 in which RNO1 from F344 is introgressed into the SHRSP backbround is indicated. |
Status of experiments for candidate gene identification by:
· consomic strain breeding – established,
· sub-congenic strain breeding – in progress,
· microarray analysis in LV tissue – in preparation,
· comparative mapping – in progress.
Experiments for candidate gene validation by:
· genetic epidemiology in MONICA study,
· functional studies in humans with LVH,
· experimental in transgenic animals,
· experimental in the zebra fish model
will start in 2006.
Topic 2:
A second goal (topic 2) of this subproject is to identify genetic pathways that confer an increased risk for the development of LV diastolic and LV systolic dysfunction in LVH. We will combine the analysis of quantitative physiological traits and genome wide quantitative expression profiling in linkage and segregation analysis (genetical genomics) in an experimental intercross [F2 (SHHF x WKY)] involving the spontaneous hypertensive heart failure (SHHF) rat to identify quantitative physiological and expression trait loci (pQTLs and eQTLs, respectively, Figure 3). Clustering of quantitative gene expression phenotypes will be used to identify co-regulated genes and to determine co-segregation of transcriptional profiles with physiological and pathophysiological phenotypes (Figure 4).
Genome-wide linkage analyses in F2-rats for physiological traits have been completed and expression profiling is currently in progress.
It is anticipated that, by defining the genetic networks and regulatory mechanisms underlying gene expression, we will identify many of the allelic variants that regulate cardiac phenotypes in this rat model for human heart failure.
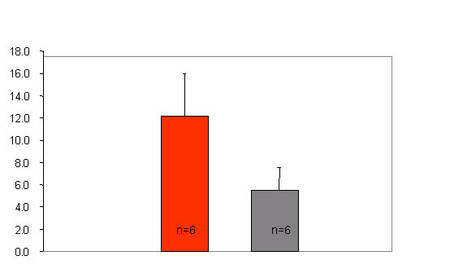 |
Fig 3: Left ventricular dysfunction in the SHHF rat. SHHF animals demonstrate significantly increased left ventricular end-diastolic pressures (LVEDP, mmHg), indicating left ventricular dysfunction in comparison to the normotensive Wistar-Kyoto rat (WKY). |
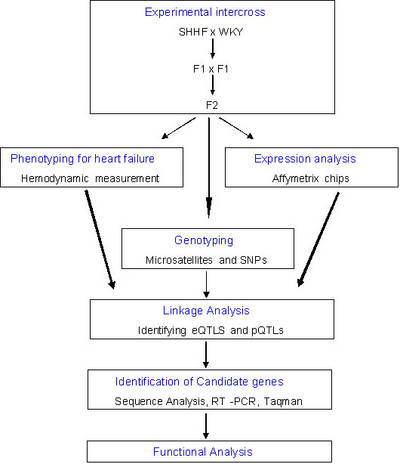 |
Fig 4: Work flow in topic 2 of KGCV1.1 |
LVH is a strong risk factor for LV dysfunction, heart failure, coronary heart disease, and arrhythmias and thus for overall cardiovascular mortality and morbidity. Thus, insights form this subproject are of mutual importance for the other research topics in the KGCV. We aim to translate genetic findings from experimental models to clinical relevance and to provide clinical investigators with target regions for genetic studies as well as model organisms for subsequent functional studies. This project represents a case in point of our networking approach established in NGFN-1 in which the added value of intensified networking can be highlighted. Here we will investigate candidate loci identified as QTL for LV mass and LV dysfunction in polygenetic rat models of hypertension in human populations. Our work flow within the NGFN is driven by the analysis of available rat models by the groups in Berlin, functional and positional candidate gene analysis via collaboraton with SMPs, comparative mapping and haplotype analysis in large population based studies characterized for LVH (MONICA study involving 1600 individuals) and controls (KORA) by our partners in Lübeck and at the SMP-GEM (GSF in München). Genotyping and statistical genetics will be performed in collaboration with the KGEV and SMP-GEM (Kiel, Münster). Subsequently, functional analysis of identified candidate genes will be carried out by extensive clinical phenotyping of patients by our partners in Erlangen, in the experimental zebra fish system by knock-down experiments in Heidelberg and the SMP-Models (GMC München)
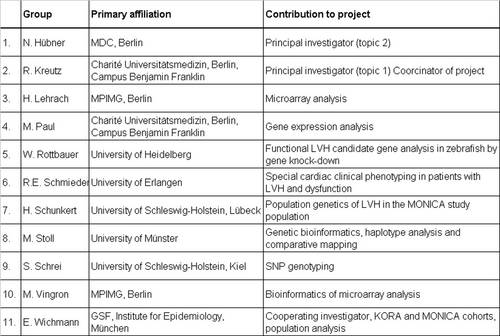 |
Table 1: List of NGFN groups participating in this network project |
Publications:
1. Kreutz R, Zurcher H, Kain S, Martus P, Offermann G, Beige J. The effect of variable CYP3A5 expression on cyclosporine dosing, blood pressure and long-term graft survival in renal transplant patients. Pharmacogenetics. 2004 Oct; 14(10):665-71.
2. Tripodi G, Florio M, Ferrandi M, Modica R, Zimdahl H, Hubner N, Ferrari P, Bianchi G. Effect of Add1 gene transfer on blood pressure in reciprocal congenic strains of Milan rats. Biochem Biophys Res Commun. 2004 Nov 12;324(2):562-8.
3. Yagil C, Hubner N, Monti J, Schulz H, Sapojnikov M, Luft FC, Ganten D, Yagil Y. Identification of hypertension-related genes through an integrated genomic-transcriptomic approach. Circ Res. 2005 Apr 1;96(6):617-25.
4. Hubner N, Wallace CA, Zimdahl H, Petretto E, Schulz H, Maciver F, Mueller M, Hummel O, Monti J, Zidek V, Musilova A, Kren V, Causton H, Game L, Born G, Schmidt S, Muller A, Cook SA, Kurtz TW, Whittaker J, Pravenec M, Aitman TJ. Integrated transcriptional profiling and linkage analysis for identification of genes underlying disease. Nat Genet. 2005 Mar;37(3):243-53.
5. Niu T, Ding AA, Kreutz R, Lindpaintner K. An expectation-maximization-likelihood-ratio test for handling missing data: application in experimental crosses. Genetics. 2005 Feb; 169(2):1021-31.
6. Kossmehl P, Schonberger J, Shakibaei M, Faramarzi S, Kurth E, Habighorst B, von Bauer R, Wehland M, Kreutz R, Infanger M, Schulze-Tanzil G, Paul M, Grimm D. Increase of fibronectin and osteopontin in porcine hearts following ischemia and reperfusion. J Mol Med. 2005 Mar 16;
7. Siegel AK et al. Genetic Loci Contribute to the Progression of Vascular and Cardiac Hypertrophy in Salt-Sensitive Spontaneous Hypertension. Arterioscler Thromb Vasc Biol Biol. 2003;23:1211-12179.
8. Kreutz R et al. The role of the cytochrome-P450 3A5 enzyme for blood pressure regulation in the general Caucasian population, Pharmacogenetics and Genomics, in press.
9. Gibbs RA et al. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature. 2004 Apr 1;428(6982):493-521.
10. Zimdahl H et al. A SNP map of the rat genome generated from cDNA sequences. Science. 2004 Feb 6;303(5659):807
more information about the working group:
http://www.charite.de/kliphatox/forschung/ag/kreutz_1.html