 » Barrier-free View
» Barrier-free View
| ||||||||
 | ||||||||
 |
 |
 |
| Picture of the 1st Network Meeting of the Herz-Kreislauf-NetzDownloadsArchivExternal job opportunities |
 |
Identification of genes for monogenic forms of myocardial infarction
 |
Subproject leader
PD Dr. Jeanette Erdmann
Department of Medicine 2
University of Schleswig-Holstein, Campus Lübeck
Ratzeburger Allee 160
23538 Lübeck
email: jeaberlin@versanet.de
Phone: 0451-500 4857
In rare cases, coronary artery disease (CAD) and myocardial infarction (MI) appear to be inherited in an autosomal-dominant mode. The identification of the underlying gene defects in these rare monogenic forms of CAD/MI will help to define genetic pathways and provide insights into molecular mechanisms for the pathogenesis of common forms of CAD and MI.
The major goal of our project is (topic 1) to identify the underlying genetic defect in families showing an autosomal-dominant inheritance pattern of MI. The identification of the genetic defect will lead to knowledge about the pathogenesis of CAD and MI and thus may be of enormous benefit for the identification of mechanisms and relevant other candidate genes. Therefore, the second goal of our project (topic 2) is to elucidate if genes identified to cause rare monogenic forms of MI are also involved in the pathogenesis of more common and possibly polygenic forms of CAD/MI.
In recent years we have identified a total of 19 multiplex families (max. 27 MI patients per family) displaying an autosomal dominant inheritance pattern using systematic large scale screening (Fig. 1).
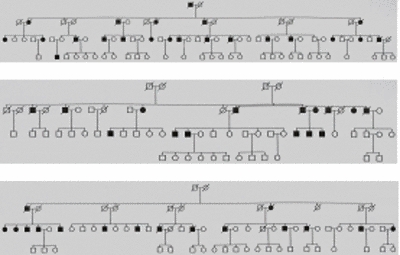 |
Fig 1: Exemplary pedigrees of our extended families with myocardial infarction. |
Power simulation using the Slink software package and applying several models of inheritance was carried out in these families. Analysis consistently revealed the highest theoretical LOD scores under the assumption of an autosomal dominant inheritance. A maximum LOD score of 3.0 was obtained in a single large family using the affected-only model. After age-adjusted consideration of the non-affected individuals, the maximum simulated LOD score increased to 6.2 and was significant in four of the 19 families.
In addition, we evaluated heritability of the binary phenotype myocardial infarction and additional quantitative phenotypes (body constitution: height, weight, and body mass index (BMI), systolic and diastolic blood pressure and lipids: cholesterol, LDL, HDL, and triglycerides) in 496 participants from 19 families.
Heritability estimates in the narrow sense were obtained by a variance-component analysis using Solar. Corresponding 95% confidence intervals and p-values were based on the one-step jack-knife approximation to the robust estimator of variance and were estimated in a script language.
Not surprisingly heritability value (h2) for myocardial infarction was particularly strong (75.3%; p<0.01), due to the selection criteria for our families of at least three or more affected living MI siblings.
Whereas heritability estimates were moderate for height (31.8%), weight (36.2%), BMI (31.5%), systolic blood pressure (29.9%), cholesterol (33.2%), LDL (28.3%), HDL (38.9%) and triglycerides (32.3%) (largest p<0.02 for all) (Table 1), these values are comparable with data from the literature.
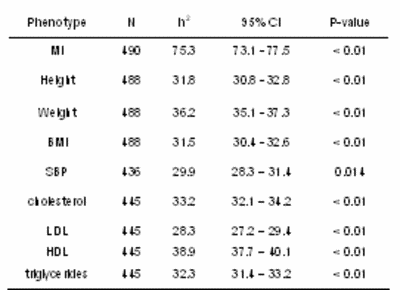 |
Tab 1: Heritability estimates h2, robust 95% confidence intervals (95% CI) and robust p-values for myocardial infarction (MI), body constitution, systolic blood pressure (SBP), and lipid parameters (HDL, LDL, and triglycerides). |
Recently, a genome-wide scan in these pedigrees was completed at the Gene Mapping Centre in Berlin, Germany (SMP-DNA) and the SMP-GEM (Bonn) (funded by DFG).
Furthermore, we have re-evaluated (set of over 700 markers, average marker distance ~ 6 cM) our most promising linkage signal on chromosome 8q24 which was revealed in our initial model-based and model-free linkage analyses using GENEHUNTER and Simwalk2 (Figure 2)
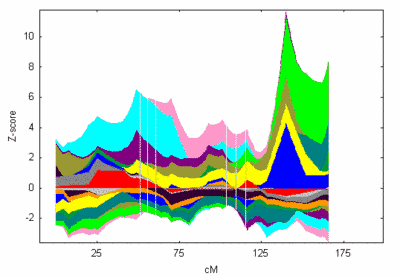 |
Fig 2: Contribution of all families to the linkage signal on chromosome 8. NPL z-all statistic as implemented in GENEHUNTER. |
This locus was identified in three unrelated families based on both, NPL score and haplotype reconstruction (figure 1 and 2). The NPL scores for three of the families are 4.3, 3.9, and 1.0, respectively. In each family, a haplotype spanning 4 markers on chromosome 8q24 between markers D8S1784 and D8S272 cosegregates with MI (see figure 2).
Within the 22-Mbp 8q24 region linked in the current families there are approximately 70 known or predicted genes, which are currently being evaluated, including two very interesting candidate genes: osteoprotegerin (TNFRSF11B, OPG) and ectonucleotide pyrophosphatase/phospodiesterase 2 (ENPP2). For example, osteoprotegerin is a secretory glycoprotein that belongs to the tumor necrosis factor receptor family. OPG-deficient mice develop severe osteoporosis and medial arterial calcification of the aorta. Furthermore OPG immunoreactivity was in early atherosclerotic lesions. Examination whether serum OPG levels are associated with the progression of coronary artery disease (CAD) revealed that serum OPG levels are associated with the presence and severity of CAD.
Recently, an genetic association study by Soufi et al. reported that genetic variations of the OPG gene may confer an increased risk of CAD in Caucasian men.
Sequence analysis is ongoing and will be finished within the framework of our project.
Recently, we systematically analyzed 906 coronary angiograms from 348 of our myocardial infarction families. After adjustment for age and gender, significant heritabilities were identified for proximal stenoses, in particular left main disease (h2=0.49±0.12; p=0.01), coronary calcification (h2=0.51±0.17; p=0.001), and ectatic coronary lesions (h2=0.52±0.07; p=0.001).
For further phenotype evaluation of our extended families, we have systematically screened the coronary morphology using angiograms which have been initiated for clinical reason. Various aspects of coronary anatomy including disease severity, distribution of lesions, presence of coronary calcifications, and morphology of stenoses have been quantified.
With respect to high rates of phenocopies as expected in multifactorial diseases, reevaluation of two point and multipoint linkage analyses in our families will reveal additional information. This setup will lead to confirmation of previous results and will generate additional putative gene loci in families with so far no positive linkage results.
The identification of genetic variants relevant for the development of rare familial CAD/MI might be also of importance for the pathogenesis of common forms of CAD/MI and might, thus, define new genetic pathways and insights into molecular mechanisms.
 |
Working group PD Dr. Erdmann |
Publications:
1. Bezzina, C. R., A. O. Verkerk, A. Busjahn, A. Jeron, J. Erdmann, T. T. Koopmann, Z. A. Bhuiyan, R. Wilders, M. M. Mannens, H. L. Tan, F. C. Luft, H. Schunkert, and A. A. Wilde. 2003. A common polymorphism in KCNH2 (HERG) hastens cardiac repolarization. Cardiovasc.Res. 59:27-36.
2. Brull, D., S. Dhamrait, S. Myerson, J. Erdmann, D. Woods, M. World, D. Pennell, S. Humphries, V. Regitz-Zagrosek, and H. Montgomery. 2001. Bradykinin B2BKR receptor polymorphism and left-ventricular growth response. Lancet 358:1155-1156.
3. Daehmlow, S., J. Erdmann, T. Knueppel, C. Gille, C. Froemmel, M. Hummel, R. Hetzer, and V. Regitz- Zagrosek. 2002. Novel mutations in sarcomeric protein genes in dilated cardiomyopathy. Biochem.Biophys.Res.Commun. 298:116-120.
4. Erdmann, J., S. Daehmlow, S. Wischke, M. Senyuva, U. Werner, J. Raible, N. Tanis, S. Dyachenko, M. Hummel, R. Hetzer, and V. Regitz-Zagrosek. 2003. Mutation spectrum in a large cohort of unrelated consecutive patients with hypertrophic cardiomyopathy. Clin.Genet. 64:339-349.
5. Erdmann, J., J. Raible, J. Maki-Abadi, M. Hummel, J. Hammann, B. Wollnik, E. Frantz, E. Fleck, R. Hetzer, and V. Regitz-Zagrosek. 2001. Spectrum of clinical phenotypes and gene variants in cardiac myosin-binding protein C mutation carriers with hypertrophic cardiomyopathy. J.Am.Coll.Cardiol. 38:322-330.
6. Fischer, M., W. Lieb, D. Marold, M. Berthold, A. Baessler, H. Lowel, H. W. Hense, C. Hengstenberg, S. Holmer, H. Schunkert, and J. Erdmann. 2004. Lack of association of a 9 bp insertion/deletion polymorphism within the bradykinin 2 receptor gene with myocardial infarction. Clin.Sci.(Lond) 107:505-511.
7. Jamshidi, Y., H. E. Montgomery, H. W. Hense, S. G. Myerson, I. P. Torra, B. Staels, M. J. World, A. Doering, J. Erdmann, C. Hengstenberg, S. E. Humphries, H. Schunkert, and D. M. Flavell. 2002. Peroxisome proliferator--activated receptor alpha gene regulates left ventricular growth in response to exercise and hypertension. Circulation 105:950-955.
8. Jeron, A., C. Hengstenberg, S. Holmer, B. Wollnik, G. A. Riegger, H. Schunkert, and J. Erdmann. 2004. KCNJ11 polymorphisms and sudden cardiac death in patients with acute myocardial infarction. J.Mol.Cell Cardiol. 36:287-293.
9. Lieb, W., R. Pavlik, J. Erdmann, B. Mayer, S. R. Holmer, M. Fischer, A. Baessler, C. Hengstenberg, H. Loewel, A. Doering, G. A. Riegger, and H. Schunkert. 2004. No association of interleukin-6 gene polymorphism (-174 G/C) with myocardial infarction or traditional cardiovascular risk factors. Int.J.Cardiol. 97:205-212.
10. Schmieder, R. E., J. Erdmann, C. Delles, J. Jacobi, E. Fleck, K. Hilgers, and V. Regitz- Zagrosek. 2001. Effect of the angiotensin II type 2-receptor gene (+1675 G/A) on left ventricular structure in humans. J.Am.Coll.Cardiol. 37:175-182.
11. Gonzalez,P. et al. The Pro279Leu variant in the transcription factor MEF2A is associated with myocardial infarction. J Med. Genet. (2005).
12. Bhagavatula,M.R. et al. Transcription factor MEF2A mutations in patients with coronary artery disease. Hum. Mol. Genet. 13, 3181-3188 (2004).
13. Wang,L., Fan,C., Topol,S.E., Topol,E.J. & Wang,Q. Mutation of MEF2A in an inherited disorder with features of coronary artery disease. Science 302, 1578-1581 (2003).
14. Coady,S.A. et al. Genetic variability of adult body mass index: a longitudinal assessment in framingham families. Obes. Res 10, 675-681 (2002).
15. Edwards,K.L., Mahaney,M.C., Motulsky,A.G. & Austin,M.A. Pleiotropic genetic effects on LDL size, plasma triglyceride, and HDL cholesterol in families. Arterioscler. Thromb. Vasc. Biol. 19, 2456-2464 (1999).
16. Fava,C. et al. Heritability of ambulatory and office blood pressure phenotypes in Swedish families. J Hypertens. 22, 1717-1721 (2004).
17. Mitchell,G.F. et al. Heritability and a genome-wide linkage scan for arterial stiffness, wave reflection, and mean arterial pressure: the Framingham Heart Study. Circulation 112, 194-199 (2005).
18. Jono,S. et al. Serum osteoprotegerin levels are associated with the presence and severity of coronary artery disease. Circulation 106, 1192-1194 (2002).
19. Soufi,M. et al. Osteoprotegerin gene polymorphisms in men with coronary artery disease. J Clin. Endocrinol. Metab 89, 3764-3768 (2004).
20. Fischer,M. et al. Distinct heritable patterns of angiographic coronary artery disease in families with myocardial infarction. Circulation 111, 855-862 (2005).
more information about the working group:
http://www.innere2.mu-luebeck.de/