 » Barrier-free View
» Barrier-free View
| ||||||||
 | ||||||||
 |
 |
 |
| Picture of the 1st Network Meeting of the Herz-Kreislauf-NetzDownloadsArchivExternal job opportunities |
 |
Genetics of coronary morphology
 |
Subproject leader
Prof. Dr. Heribert Schunkert
Department of Medicine 2
University of Schleswig-Holstein,
Campus Lübeck
Ratzeburger Allee 160
23538 Lübeck
email: heribert.schunkert@innere2.uni-luebeck.de
Phone: 0451-500 2501
Myocardial infarction (MI) is both multifactorial in origin and phenotypically diverse. Multiple etiologic components and multiple morphological features of CAD make it likely that different causes lead to different disease patterns. Thus, in addition to differentiate factors leading to CAD, it is crucial to precisely specify distinct features of coronary morphology. Accordingly, we hypothesize intermediate phenotypes reflecting certain aspects of CAD facilitate the identification of distinct gene-phenotype relationships. Particularly, the phenotypic characterization of the coronary morphology may enable to specifically investigate whether distinct CAD patterns reflect distinct genetic causes. The importance of such substratification has recently been documented in the identification of the first stroke-related gene.
In order to overcome the complexity of CAD in humans, we will also study the phenotype in animal models. In fact, man, mouse and rat share common features of cardiovascular calcification. Recently, we identified two significant QTLs (Dyscalc1 and Dyscalc2) linked to dystrophic cardiac calcification. Moreover, we mapped a QTL suggestive for linkage with coronary calcification in a human region syntenic to Dyscalc2, providing a region of high priority to follow up in human studies.
Studies in humans heritability of coronary phenotypes
We retrospectively studied the coronary angiograms of 882 siblings with CAD from 401 families (Figure 1). These families were ascertained through index patients defined by MI before the age of 60 years and at least one sibling with MI or coronary revascularization procedures. Heritability calculations were performed using variance component analysis. Additionally, recurrence risks to siblings were analyzed.
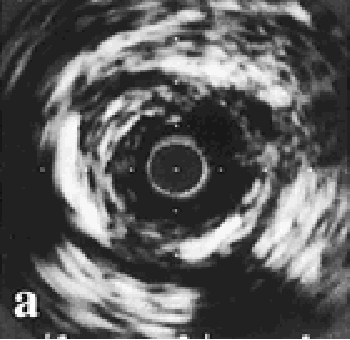 |
Fig 1: Coronary calcification. |
Traditional cardiovascular risk factors and age at the first coronary event displayed significant heritable components. After adjustment for age and gender, significant heritabilities were identified for proximal stenoses, in particular left main disease (h2=0.49±0.12; p=0.01), coronary calcification (h2=0.51±0.17; p=0.001), and ectatic coronary lesions (h2=0.52±0.07; p=0.001) (Figure 2).
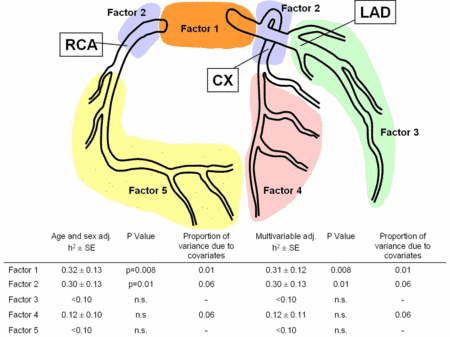 |
Fig 2: Schematic overview of the factors resulting from principal component factor analysis with their heritability values. |
In contrast, no heritability was found for distal disease (h2=0.05±0.19; n.s.), the pattern of coronary arterial blood supply or the number of diseased vessels (Fig. 2).
To confirm the heritability measures we analysed a second set of 1,500 coronary angiograms from MI families and tested for heritability of selected coronary phenotypes.
The analyses are ongoing; but preliminary results confirm the heritability measures in this second sample.
Animal studies unravelling the Dyscalc1 locus
Dyscalc1 was mapped to a chromosomal region on mouse proximal chromosome 7. Since, Dyscalc1 was intensively investigated. We localized Dyscalc1 to a 10 cM chromosomal segment between D7Mit227 and D7Mit230 microsatellite markers. We confirmed the contribution of Dyscalc1 to dystrophic cardiac calcification (DCC) using a congenic line that carry the DCC-susceptible allele on a resistant genetic background. We demonstrated that Dyscalc1 contribute not only to cardiac calcification but also to aorta calcification. Furthermore, using bone marrow transplantation, we showed that calcium phosphate deposit was initiated in DCC-susceptible strains but influenced by infiltrating bone marrow cells.
The aim of the ongoing project is to identify, among the 160 genes located in Dyscalc1 critical region, putative candidate genes representing the so far unknown Dyscalc1 gene(s).
Differential gene expression
On the basis of differential gene expression patterns, which were determined in various tissues from resistant C57BL/6 and susceptible C3H/He mice, we identified putative candidate genes in the Dyscalc1 critical region. Specific primer pairs for each gene of the 160 genes located in Dyscalc1 interval were designed and polymerase chain reaction (PCR) has been performed for detecting the gene expression in different organ tissue. Genes that were expressed in myocardial tissue have been further investigated using Real-time RT-PCR (Figure 3).
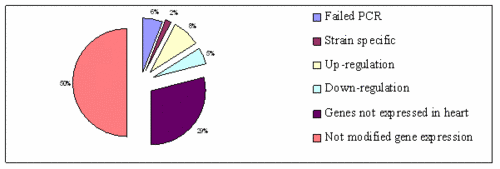 |
Fig 3: Differential gene expression analysis using real-time RT-PCR for genes located in the Dycsalc1 critical region |
Sequence analyses of putative candidate genes are ongoing and experiments are underway to further examine the biological functions of genetic variants in these candidate genes in mice and humans.
In-silico mapping of Dyscalc1 critical region
In a complementary approach we further tried to narrow down the Dyscalc1 critical region using in silico-mapping.
Analyzing recombination among eight laboratory strains within the Dyscalc1 segment further narrowed down this region to a 1-Mb segment within an in silico QTL peak with a significant R value of 1.8 (Figure 4). These results demonstrate that in silico-mapping is a powerful tool that could be used to confirm fine mapping results of classical QTL analysis.
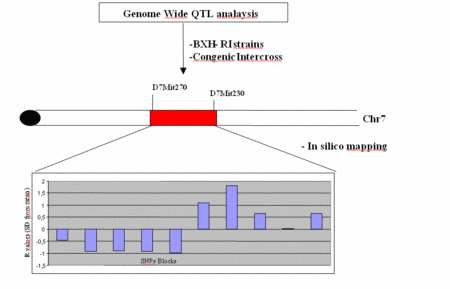 |
Fig 4: Schematic presentation of the in silico mapping approach and results. |
Human Studies Identification of genetic variants in candidate genes and association studies
Until now, we sequenced five candidate genes (5´UTR, 3´UTR, promoter, and coding exons) within the Dyscalc1 critical region in a subgroup of 48 patients with different degrees of coronary calcification.
The frequency of variants leading to amino acid substitutions will be further determined in larger sets of patients with different coronary phenotypes and controls. Our aim is to identify genetic variants associated with specific coronary phenotypes.
Publications:
1. Bezzina, C. R., A. O. Verkerk, A. Busjahn, A. Jeron, J. Erdmann, T. T. Koopmann, Z. A. Bhuiyan, R. Wilders, M. M. Mannens, H. L. Tan, F. C. Luft, H. Schunkert, and A. A. Wilde. 2003. A common polymorphism in KCNH2 (HERG) hastens cardiac repolarization. Cardiovasc.Res. 59:27-36.
2. Broeckel, U., C. Hengstenberg, B. Mayer, S. Holmer, L. J. Martin, A. G. Comuzzie, J. Blangero, P. Nurnberg, A. Reis, G. A. Riegger, H. J. Jacob, and H. Schunkert. 2002. A comprehensive linkage analysis for myocardial infarction and its related risk factors. Nat.Genet. 30:210-214.
3. Hengstenberg, C., S. R. Holmer, B. Mayer, H. Lowel, S. Engel, H. W. Hense, G. A. Riegger, and H. Schunkert. 2000. Evaluation of the aldosterone synthase (CYP11B2) gene polymorphism in patients with myocardial infarction. Hypertension 35:704-709.
4. Holmer, S. R., C. Hengstenberg, H. G. Kraft, B. Mayer, M. Poll, S. Kurzinger, M. Fischer, H. Lowel, G. Klein, G. A. Riegger, and H. Schunkert. 2003. Association of polymorphisms of the apolipoprotein(a) gene with lipoprotein(a) levels and myocardial infarction. Circulation 107:696-701.
5. Jamshidi, Y., H. E. Montgomery, H. W. Hense, S. G. Myerson, I. P. Torra, B. Staels, M. J. World, A. Doering, J. Erdmann, C. Hengstenberg, S. E. Humphries, H. Schunkert, and D. M. Flavell. 2002. Peroxisome proliferator-- activated receptor alpha gene regulates left ventricular growth response to exercise and hypertension. Circulation 105:950-955.
6. Jeron, A., C. Hengstenberg, S. Holmer, B. Wollnik, G. A. Riegger, H. Schunkert, and J. Erdmann. 2004. KCNJ11 polymorphisms and sudden cardiac death in patients with acute myocardial infarction. J.Mol.Cell Cardiol. 36:287-293.
7. Pokharel, S., P. P. van Geel, U. C. Sharma, J. P. Cleutjens, H. Bohnemeier, X. L. Tian, H. Schunkert, H. J. Crijns, M. Paul, and Y. M. Pinto. 2004. Increased myocardial collagen content in transgenic rats overexpressing cardiac angiotensin-converting enzyme is related to enhanced breakdown of N-acetyl-Ser-Asp-Lys-Pro and increased phosphorylation of Smad2/3. Circulation 110:3129-3135.
8. Schunkert, H., U. Brockel, C. Hengstenberg, A. Luchner, M. W. Muscholl, K. Kurzidim, B. Kuch, A. Doring, G. A. Riegger, and H. W. Hense. 1999. Familial predisposition of left ventricular hypertrophy. J.Am.Coll.Cardiol. 33:1685-1691.
9. Schunkert, H., H. W. Hense, S. R. Holmer, M. Stender, S. Perz, U. Keil, B. H. Lorell, and G. A. Riegger. 1994. Association between a deletion polymorphism of the angiotensin-converting-enzyme gene and left ventricular hypertrophy. N.Engl.J.Med. 330:1634-1638.
10. Sedlacek, K., M. Fischer, J. Erdmann, C. Hengstenberg, S. Holmer, S. Kurzinger, M. Muscholl, A. Luchner, G. A. Riegger, H. W. Hense, and H. Schunkert. 2002. Relation of the G protein beta3-subunit polymorphism with left ventricle structure and function. Hypertension 40:162-167.
11. Lieb W, Graf J, Gotz A, Konig IR, Mayer B, Fischer M, Stritzke J, Hengstenberg C, Holmer SR, Doring A, Lowel H, Schunkert H, Erdmann J. Association of angiotensin-converting enzyme 2 (ACE2) gene polymorphisms with parameters of left ventricular hypertrophy in men Results of the MONICA Augsburg echocardiographic substudy. J Mol Med. 2005 Nov 11; [Epub ahead of print] PMID: 16283142 [PubMed - as supplied by publisher]
12. Mayer B, Lieb W, Gotz A, Konig IR, Aherrahrou Z, Thiemig A, Holmer S, Hengstenberg C, Doering A, Loewel H, Hense HW, Schunkert H, Erdmann J. Association of the T8590C polymorphism of CYP4A11 with hypertension in the MONICA Augsburg echocardiographic substudy. Hypertension. 2005 Oct; 46(4):766-71. Epub 2005 Sep 6. PMID: 16144986 [PubMed - indexed for MEDLINE]
13. Lieb W, Pavlik R, Erdmann J, Mayer B, Holmer SR, Fischer M, Baessler A, Hengstenberg C, Loewel H, Doering A, Riegger GA, Schunkert H. No association of interleukin-6 gene polymorphism (-174 G/C) with myocardial infarction or traditional cardiovascular risk factors. Int J Cardiol. 2004 Nov; 97(2):205-12. PMID: 15458685 [PubMed - indexed for MEDLINE]
14. Fischer M, Lieb W, Marold D, Berthold M, Baessler A, Lowel H, Hense HW, Hengstenberg C, Holmer S, Schunkert H, Erdmann J. Lack of association of a 9 bp insertion/deletion polymorphism within the bradykinin 2 receptor gene with myocardial infarction. Clin Sci (Lond). 2004 Nov;107(5):505-11. PMID: 15301669 [PubMed - indexed for MEDLINE]
15. Doehring LC, Kaczmarek PM, Ehlers EM, Mayer B, Erdmann J, Schunkert H, Aherrahrou Z. Arterial calcification in mice after freeze-thaw injury. Annals of Anatomy (im Druck)
16. Gretarsdottir S et al. (2003) The gene encoding phosphodiesterase 4D confers risk of ischemic stroke. Nat Genet. 35:131-8.
17. Ivandic BT et al. (1996) A locus on chromosome 7 determines myocardial cell necrosis and calcification (dystrophic cardiac calcinosis) in mice. Proc Natl Acad Sci U S A. 93:5483-8.
18. Ivandic BT et al. (2001) New Dyscalc loci for myocardial cell necrosis and calcification (dystrophic cardiac calcinosis) in mice. Physiol. Genomics 6:137-44.
19. Fischer M. et al. (2005) Distinct heritable patterns of angiographic coronary artery disease in families with myocardial infarction. Circulation 111, 855-862.
20. Aherrahrou et al. (2004) A locus on chromosome 7 determines dramatic up-regulation of osteopontin in dystrophic cardiac calcification in mice. Am J Pathol. 164:1379-87.
21. Kazmarek et al. (2005) Reduced cardiac calcinosis in DCC-susceptible congenic mice reconstituted with DCC-resistant bone marrow cells. Poster presentation; European Society of Cardiology, Stockholm, Sweden.
22. Doehring et al. (2005) Dyscalc1 determines aortic calcification in mice. Poster presentation; European Society of Cardiology, Stockholm, Sweden.
23. Grupe A et al. (2001) In silico mapping of complex disease-related traits in mice Science. 292(5523): 1915-8.
more information about the working group: