 » Barrier-free View
» Barrier-free View
| ||||||||
 | ||||||||
 |
 |
 |
| Picture of the 1st Network Meeting of the Herz-Kreislauf-NetzDownloadsArchivExternal job opportunities |
 |
Contribution of genetic variation in modifier genes to cardiomyopathy phenotypes
 |
Subproject leader
Prof. Dr. Hugo A. Katus
University of Heidelberg
Internal Medicine III / Cardiology and Pulmonary Medicine
Im Neuenheimer Feld 410
69120 Heidelberg
email: sekretariat_katus@med.uni-heidelberg.de
Phone: 06221-56 8670
cardiomyopathy (DCM) phenotype may vary considerably among family members carrying identical mutations. Common genetic variation in modifier genes and ethnic background contribute to the observed variability. Since inflammation is linked to the progression of contractile dysfunction, we genotyped a well characterized study cohort of 792 patients with contractile dysfunction and 721 controls for a variety of genes involved in inflammation. These studies revealed a significant association with polymorphisms in the IL1-, IL6-, IL8-, IL10- and in other pathways, such as the Toll-like receptor signalling pathway (Fig. 1). Now, the goal of the study is to confirm and extend these findings by typing additional SNPs in genes linked to inflammation in an extended cohort of 1500 DCM patients and 2000 controls. This analysis will use significantly associated SNPs, which were identified in NGFN-1, and additional SNPs, which were characterized by the NGFN network of environmental diseases in studies of inflammatory bowel disease. Then, the contribution of potential DCM modifier genes derived from both studies, will be also assessed in a cohort of 150 cases and 300 controls of African ethnicity. This will facilitate the characterization and resolution of potential haplotype blocks, which are expected to be considerably smaller in this phylogenetically older population. Finally, molecular and cell biological studies will ensue to understand the functional consequences of the relevant genetic modifiers.
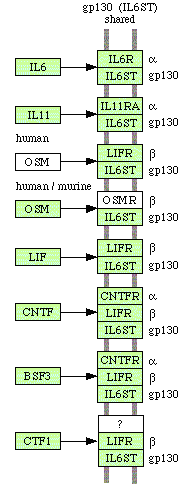 |
Fig.1 Pathways involving Interleukin 6 and related proteins (from the KEGG database kegg.com/dbget-bin/get_pathway). |
DCM patients study cohort. For genetic analyses of inflammatory genes we use a collection of approximately 1500 DCM patients recruited by H.A. Katus (792 patients),
V. Regitz-Zagrosek, German Heart Centre Berlin (528 patients) and 350 patients expected from PopGen. Directed by Prof. S. Schreiber and Prof. M. Krawczak, University of Kiel in Germany, PopGen is a NGFN research initiative, which implemented broad phenotyping of patients covering all common disorders including inflammatory diseases. The population of Northern Germany was included in this comprehensive assessment of health and several thousand of control subjects were recruited for population-based association studies. We use 2000 of these age-matched controls for our studies.
SNP Genotyping. Large scale SNP genotyping is performed at the SMP-DNA Kiel (directed by Prof. S. Schreiber) using multiplex TaqMan technology. Small scale genotyping, i.e. genotyping of the African cohort (contributed by D. Hilfiker, University of Hannover) or specific candidate genes, is done at in Muenster in collaboration with Prof. M. Stoll (University of Muenster, Germany).
Genetic Analyses. The primary genetic analyses are performed by the group of M. Stoll, supported by M. Krawczak (University of Kiel) and M. J. Daly (Whitehead/MIT Centre for Genome Research, Cambridge, MA U.S.A.). Each marker is tested for Hardy-Weinberg equilibrium in the control populations using a c² test. A case/control analysis will be performed with unrelated, matched controls and the German and African study cohorts. Genotype-based odds ratios (OR) are calculated for both, single markers and haplotypes. The expectation maximisation (EM) algorithm is used to estimate haplotype frequencies and pair-wise LD. The haplotype frequencies are validated with data from the HapMap Consortium using the Haploview software package provided by M. J. Daly. Gene-gene interactions are tested by logistic regression using the genotype at a given SNP marker/gene locus as covariate and also a novel algorithm, which has been applied for the analysis of inflammatory bowel disease.
A subset of patients was phenotyped for blood levels of inflammatory chemo- and cytokines. Preliminary analysis revealed a significant correlation of IL6 serum concentrations with indexes of contractile function such as ejection fraction (p=0.045) and pulmonary artery pressure (p=0.038). In NGFN-1, we genotyped a well characterized study cohort of 792 patients with contractile dysfunction and 721 controls for a variety of genes involved in inflammation (Table 1).
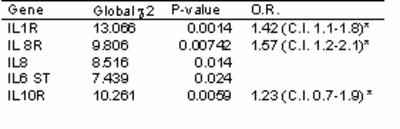 |
Table 1. Association of selected SNPs in the Interleukin pathways with DCM in 792 patients vs. 721 control individuals (significance thresholds and 95% confidence intervals were determined by permutation analysis). |
Altogether, we have now typed these and other SNPs in over 1800 patients and 3000 controls. First preliminary analysis suggested robust associations of some of the earlier results in all cohorts examined so far. At present, various models of inheritance and epistasis are calculated to reveal allelic and gene interaction. Additional genes, which are functionally linked with already positively associated genes and pathways, are selected and typed to support our encouraging, preliminary results.
The identification of modifier genes, eg. the contribution of a pro-inflammatory background to the progression of DCM, will provide valuable insights into the aetiology of contractile dysfunction and may help to improve diagnosis, prognosis and therapy of the disease. These modifier genes will provide new targets for the study in animal models and functional analyses. The close interaction with PopGen and other disease networks will facilitate the translation of our study results to other diseases and the evaluation on a population-based level across classic organ-centered disease definitions.
In cooperation with:
Prof. Dr. Stoll, GEM, Westfälische-Wilhelms-University of Münster,
Prof. Dr. Regitz-Zagrosek, German Heart Center, Berlin,
Dr. C. Zugck, University of Heidelberg, Internal Medicine III / Cardiology,
Dr. N. El Mokhtari, popgen Study Group,
University of Schleswig-Holstein, Campus Kiel,
Dr. M. Müller-Bardorff und J. Boos, University of Schleswig-Holstein, Campus Lübeck.
Publications:
1. Brull D, et al. Bradykinin B2BKR receptor polymorphism and left-ventricular growth response. Lancet. 2001; 358:1155-6
2. Erdmann J, et al. Spectrum of clinical phenotypes and gene variants in cardiac myosin-binding protein C mutation carriers with hypertrophic cardiomyopathy. J Am Coll Cardiol. 2001; 38:322-30
3. Schmieder RE, et al. Effect of the angiotensin II type 2-receptor gene (+1675 G/A) on left ventricular structure in humans. J Am Coll Cardiol. 2001; 37:175-82
4. Regitz-Zagrosek V, et al. Novel mutation in the alpha-tropomyosin gene and transition from hypertrophic to hypocontractile dilated cardiomyopathy. Circulation. 2000;102:E112-6
5. Grunig E, Tasman JA, Kucherer H, Franz W, Kubler W, Katus HA. Frequency and phenotypes of familial dilated cardiomyopathy. J Am Coll Cardiol. 1998;31: 186-94.
6. Franz WM, Muller OJ, Katus HA. Cardiomyopathies: from genetics to the prospect of treatment. Lancet. 2001;358: 1627-37.
7. Giallourakis C, Stoll M, Miller K, Hampe J, Lander ES, Daly MJ, Schreiber S, Rioux JD. IBD5 is a general risk factor for inflammatory bowel disease: replication of association with Crohn disease and identification of a novel association with ulcerative colitis. Am J Hum Genet. 2003;73:205-11.
8. Reich DE, Cargill M, Bolk S, Ireland J, Sabeti PC, Richter DJ, Lavery T, Kouyoumjian R, Farhadian SF, Ward R, Lander ES. Linkage disequilibrium in the human genome. Nature. 200;411:199-204.
more information about the working group:
http://www.med.uni-heidelberg.de/med/med3/index.html